Аннотация
Beer DNA authentication is the process of authentication by identification of barley malt Hordeum vulgare or its substitutes, as well as hops and yeast. The method is based on molecular genetic analysis of residual quantities of nucleic acids extracted from the cellular debris of the final product. The aim of the study was to analyse scientific and methodical approaches to extraction of residual quantities of beer raw materials nucleic acids and beer DNA authentication for their later application in determining brewing products authenticity. The technological level discloses the method of DNA extraction from wines, modified for extraction of nucleic acids from beer samples. The method includes the following characteristic peculiarities: stage enzymatic hydrolysis of polysaccharides and polypeptides of dissolved lyophilisate, multiple sedimentation and resursuspension of nucleoproteid complex, RNA removal followed by DNA extraction by organic solvents, and additional DNA purification by magnetic particle adsorption. This review presents the analysis of genetic targets used as molecular markers for gene identification of malting barley varieties and beer DNA authentication. We also provided the interpretation of PCR analysis of Hordeum vulgare varieties and samples of commercial beer. Data on SSR- and SNP-markers of Hordeum vulgare nuclear DNA, used for barley varieties identification and potentially suitable for beer DNA authentication, are also presented. We also analysed genetic targets used in malting barley substitute detection, as well as hops and yeast identification in beer. Data on correlation of amplified DNA targets with beer quality indicators were systematised.Ключевые слова
Alcoholic beverages, malting barley, Hordeum vulgare, DNA, authentication, identification, marker, PCRВВЕДЕНИЕ
Wide assortment of brewery products and their multicomponent composition refers them to the segment of difficult-to-identify goods. Their authentication is aimed at protecting consumers and manufacturers’ rights [1].
One of the strategically important tasks achievable by multidisciplinary science-intensive approaches is the search for objective identification criteria with a high degree of authenticity assessment of brewery products [2].
Molecular and genetic research methods can provide the technological process of DNA authentication of beer brands [3], thereby expanding the complex scheme of brewery products identification, traditionally based on documentary, visual, sensory and physical and chemical analyses [4].
Beer brands DNA authentication is a technological process of the authenticity verification by the gene identification of Hordeum vulgare barley malt, or its substitutes, as well as its key ingredients – hops and yeast, by molecular genetic analysis of residual quantities of nucleic acids extracted from the cellular debris of the products [3].
The analysis of scientific and methodological approaches points to the applicability of DNA technologies for detecting counterfeit and falsified brewery products.
РЕЗУЛЬТАТЫ И ИХ ОБСУЖДЕНИЕ
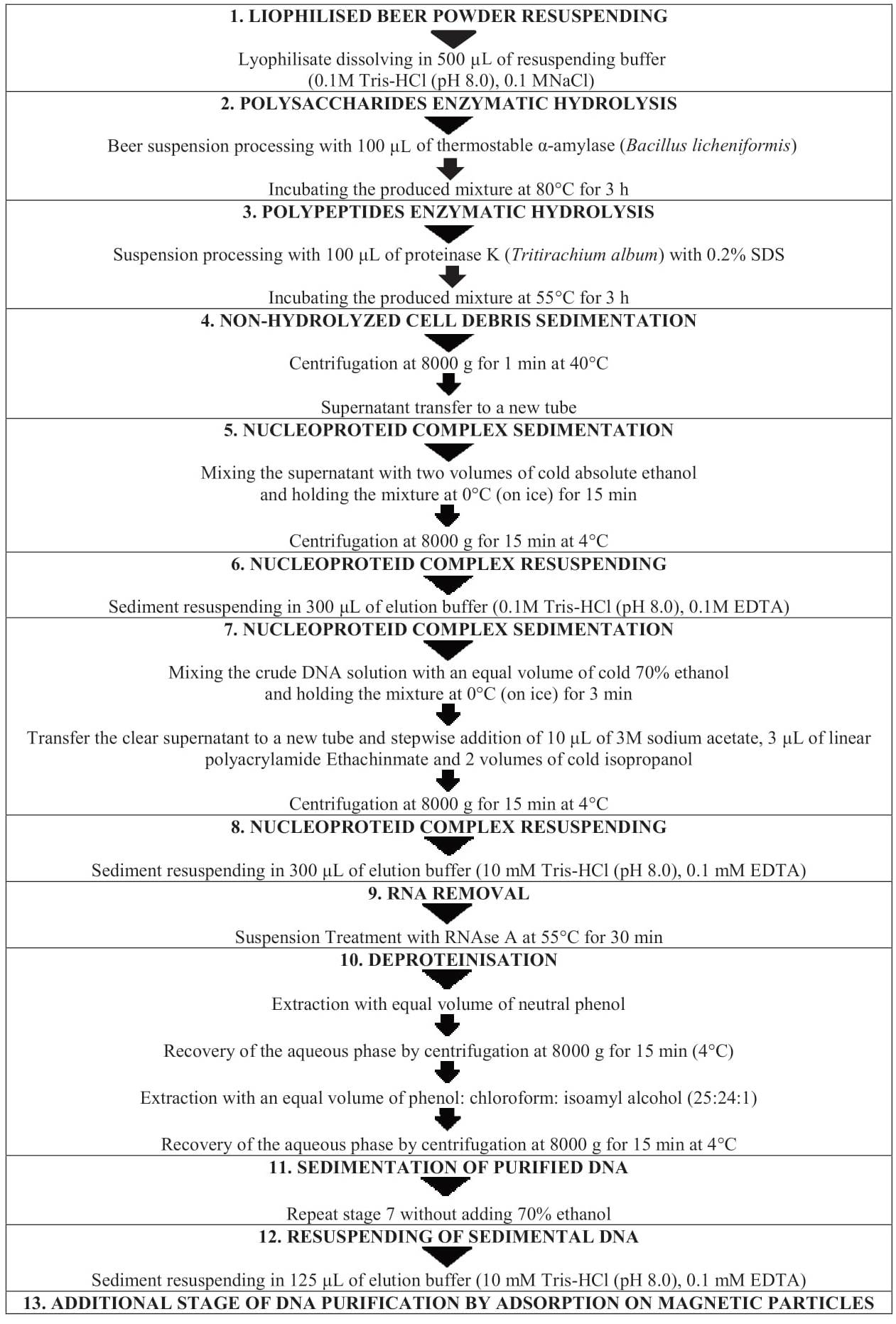
Extraction of DNA residues of beer raw materials. The technological level discloses a method for DNA extraction from wines [5, 6]. It was later modified for extraction of nucleic acids from beer samples [3]. The method includes the following characteristic peculiarities: stage enzymatic hydrolysis of polysaccharides and polypeptides of dissolved lyophilisate, multiple sedimentation and resursuspension of nucleoproteid complex, RNA removal followed by DNA extraction by organic solvents, and additional DNA purification by magnetic particle adsorption.
Figure 1 demonstrates stages of DNA extraction according to the modified method. In particular, enzymatic hydrolysis of polysaccharides by α-amylase (Bacillus licheniformis) takes 3 h instead of 1 h, when DNA is extracted from wines [3, 5]. The time of enzymatic hydrolysis of polypeptides by proteinase K (Tritirachium album) is also increased up to 3 h. The sedimentation time of non-hydrolysed cellular debris by centrifugation at 8000 g is reduced to 1 min instead of 15 min when DNA is extracted from wines. At the stage of DNA extraction from the lyophilised beer powder, the sedimentation of the nucleoprotein complex is carried out by mixing the supernatant with two volumes of cold absolute ethanol instead of two volumes of cold isopropanol. At the next stage we mixed a solution of unpurified DNA with an equal volume of 70% ethanol. The maturing of the mixture at 0°C takes 3 min instead of 10 min, as with wines. During the subsequent nucleoprotein complex sedimentation, along with the stepwise addition of 10 μL of 3M sodium acetate and two volumes of cold isopropanol to the pre-transferred transparent supernatant, 3 μL of Ethachinmate linear polyacrylamide is added. After RNA removal and deproteinisation, the sedimentation of purified DNA is carried out without adding 70% ethanol. (Сf. DNA extraction from wines involves in the nucleic acids sedimentation in 0.2 M NaCl and two volumes of cold ethanol, followed by washing with 70% ethanol). Later, nucleic acids precipitate, resuspended in the elution buffer, undergoes an additional purification by adsorption on magnetic particles, which is one of the key modification elements of the method for extracting residual DNA of beer raw materials [3].
The ability of magnetic particles to bind DNA reversibly and easily be deposited from the suspension in the magnetic field ensures high quality of nucleic acids purification and their preservation. Magnetic particles, as a rule, are a paramagnetic core with a highly developed surface covered with a polymer film with exposed covalent-bond carboxylic groups. Magnetic tripods, used in manual and automated modes, are made of neodymium magnets resistant to demagnetisation.
The additional purification by adsorption on magnetic particles of the modified method of extraction of nucleic acids from beer samples actually took the place of polymer polyvinylpyrrolidone widely used to reduce the inhibitory effect of polyphenols on PCR [3, 7–10].
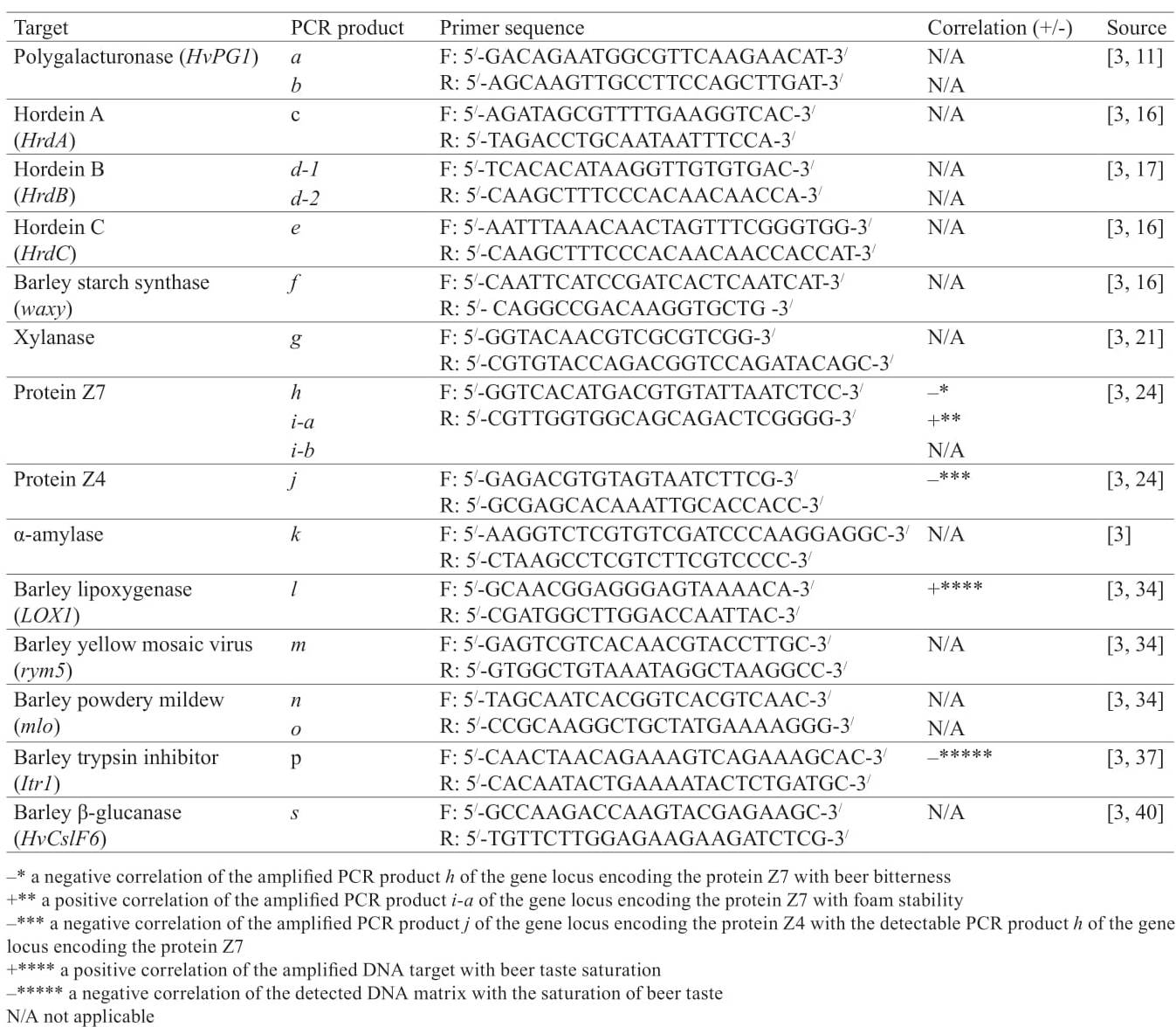
Approaches to beer DNA authentication. Genetic targets, used as molecular markers for malting barley varieties identification, can also be analysed for commercial beer DNA authentication (Table 1) [3].
Polygalacturonase is an enzyme that performs hydrolytic cleavage of α-1,4-glycoside bonds in pectin. The DNA target was the locus of its gene (HvPG1) eamplified by a corresponding pair of primers constructed by Pulido et al. based on the analysis of expressed sequence tag (EST) deposited in GenBank (A/N: EF427919) [11]. The generated PCR products a and b of the HvPG1 gene locus detected in the barley and beer samples were 89% and 79% identical to the previously deposited nucleotide sequence mRNA polygalacturonase Hordeum vulgare. Among the studied Japanese barley varieties, only the high quality ‘Ryofu’, recommended for brewing, generated two discrete fragments (a, b), like most American and Australian barley varieties, except for Stimling (Table 2). All the beer samples were marked only by the country of manufacture. They generated the PCR product b and more than half of the samples generated the additional fragment a (Table 2). The analysed DNA target was included in the group of DNA markers of identification and differentiation of beer samples, but did not correlate with the indicators of beer quality [3].
Hordeins are polymorphic proteins of barley grain coded by 7 HrdA-G loci which are localised in the short arm of the 5th Hordeum vulgare chromosome [12, 13]. Due to the established connection of the hordein-coding loci alleles with brewing qualities of barley grain, this block of targets is a priority for molecular and genetic analysis [14, 15]. From the three analysed loci (HrdA, HrdB and HrdC) only one (HrdC) was able to identify a single sample of beer out of 22 investigated by the presence of a specific PCR product e (Table 2) [3]. However, high variability of HrdA locus (up to 90% identity of nucleotide sequences of compared barley varieties with corresponding reference sequence (GenBank A/N: AF474373) indicates a certain potential of DNA authentication of beer on the analysed target by sequencing the amplified locus. The block of DNA targets under study also did not correlate with the indicators of beer quality [3].
Amylosis content in barley starch influences the quality of malt barley. Therefore, waxy-barley varieties may be a preferred option for their malting in brewing because starch with low amylosis content is more susceptible to enzymatic hydrolysis [18]. Molecular mechanism is embedded in Hordeum vulgare waxygenes located on 7 HS chromosome. They lead to the elimination of granule-bound starch synthase (GBSS) [18, 19]. Primers selected for Waxy-locus amplification had the positive control status due to generation of specific PCR product in all the samples of barley and beer [3]. Their sequeneed nucleotide DNA sequences were identical to each other and showed 98% identity to the corresponding reference Hordeum vulgare subsp. Vulgare sequence, previously deposited to GenBank (A/N: X07931) [20].
Hemicelluloses are vegetable homo- and heteropolysaccharides, which are an integral part of the endosperm cell walls. The highest content of xylanes was reported to be among the main components of hemicellulose [21]. The malt barley softens as a result of the decomposition of the cell wall. Xylanase is involved in the degradation of xylanes to xylooligosarachides, whose gene locus was used as a target for primers originally designed for DNA analysis of rice samples [3]. It is noteworthy that among the 16 varieties of barley, only three varieties (Metcalfe, Nishinohoshi and Ryofu) showed a positive amplification signal (Table 2). At the same time, due to possible obtaining inconclusive data, the authors [3] presented neither the results of PCR of beer samples, nor data on amplification of the HrdB locus.
Barley Z proteins are the main beer protein which influence beer quality, especially foam stability [22–24]. In addition, Z4 and Z7 proteins can be used as positive and negative markers of foam stability [25]. DNA-markers of foam stability developed by Limure et al. were also used in by Nakamura et al. for barley varieties identification and beer DNA-authentication [3, 25]. Identifying and differentiating barley and beer samples procedure by the gene locus, encoding proteins Z4 a nd Z7 d iffer. I n t he fi rst c ase P CR analysis is performed by interpreting three discrete PCR products (h, i-a, i-b), and in the second –by the presence or absence of a specific fragment j.
Based on the analysis, the authors recommended the further use of the tested primers for amplification of the analysed gene loci [3]. In addition, a negative correlation of the amplified PCR product h gene locus encoding Z7 protein with beer bitterness, as well as a positive correlation of PCR product i-a similar locus with foam stability (Table 1) were revealed.
Many enzymes, incl. α-amylase and β-amylase, are activated in the malting process [26, 27]. Their substrates are amylosis and amylopectin or products of their hydrolysis. Primers developed on the basis of nucleotide sequence of the gene locus encoding α-amylase initiated the amplification of PCR product k in most of the barley varieties and beer samples (Table 2) [3, 28]. It is noteworthy that the amino acid sequence of the target had 69% identity with Mla-locus of resistance to powdery mildew Hordeum vulgare (GenBank A/N: AF427791) [29]. The used set of primers was included in the group of molecular labeling systems of barley varieties, and therefore has a certain potential of practical application for beer authentication, although the authors did not mention it [3].
Lipoxygenase-deficient barley varieties with reduced or lost activity of LOX genes have a positive impact on quality indicators such as beer taste and foam stability [30–33]. The set of primers constructed by Nagamine et al. resulted in amplification of the specific PCR product l in a small number of studied barley varieties and in more than half of beer samples, whose sequenced nucleotide sequences had 99% identity with the reference sequence of locus LoxA-gene Hordeum vulgare (GenBank A/N: L35931) [3, 34, 35]. The tested set of primers was recommended for further use in the amplification of the analysed gene locus for barley varieties identification and beer brands differentiation. It should also be noted that the authors [3] additionally revealed a positive correlation between the amplified DNA target and beer taste saturation (Table 1).
The selection of barley varieties with genetic resistance to viral, bacterial and fungal diseases is aimed at high-quality grain production [36]. A number of DNA markers of resistance of barley to yellow mosaic virus (rym5-locus) and powdery mildew (mlo-locus) [34] integrated into breeding programs can also be used in molecular labelling of brewing barley varieties, which is clearly demonstrated in the work [3]. The authors interpreted the PCR analysis data of barley samples taking into account the presence or absence of specific PCR products m (rym), n and o (mlo) recorded on the corresponding electrophoregrams. But the results of the PCR analysis of beer samples and their correlation with quality indicators were not provided [3].
Protein inhibitors of proteolytic enzymes play an important role both in formation of homeostatic reactions in plants and in the process of seed maturation and germination. Selected primers to the trypsin inhibitor (Itr1) gene locus led to the amplification of the specific PCR product p in half of the tested barley varieties and beer samples [37]. Thus, the DNA marker was concluded to be highly informative [3]. Additionally, the DNA sequences of the Itr1-gene locus of the material had 94% identity with the same locus of the Hordeum vulgare subsp. vulgare gene (GenBank A/N: (X65875) [38]. Also, in the study [3] a negative correlation of the detected DNA matrix with beer taste saturation was revealed (Table 1).
The content (1–3, 1–4) of β-D-glucan in barley grain, which determines its hardness, is much higher compared to other cereals [39]. However, for barley varieties used in brewing, a lower the content of this polysaccharide in the grain is desirable in order to achieve a more effective flow of the malting process [40]. The amplification procedure of the locus of the HvCslF6 gene with a selected primer pair led to the production of a specific PCR product s in a number of American, Australian and Japanese brewing barley varieties [3, 40]. The most of the beer samples also gave a positive amplification signal (Table 2). The obtained amino acid sequence of the target had 83% identity with Hordeum vulgare CslF6-gene (GenBank A/N: EU267181) [41]. The used primer set was also included in the group of systems of barley varieties molecular labelling and beer DNAauthentication [3].
Microsatellites are widely used molecular markers which are suitable for identification of Hordeum vulgare. A wide variety of SSR-markers are being used [42–44]. Tomka et al. described a high potential of the five SSRmarkers for brewing barley varieties identification [45].
Table 3 shows the sequence of oligonucleotide primers of the corresponding Hordeum vulgare SSRmarkers of nuclear DNA, as well as the range of lengths of detected alleles and their number. The genetic identification procedure includes PCR method with subsequent data interpretation by horizontal or vertical gel electrophoresis and DNA fragmentary analysis of capillary gel electrophoresis. The SSR-markers, potentially suitable for beer DNA authentication, are advisable to test in the formulation of single PCR, with a set of primers of a single SSR-marker to achieve a reproducible result.
Alongside with SSR-markers, SNP-markers, used for barley varieties identification, including brewing ones, also have high identification capacity [46–48].
Table 4 shows oligonucleotide primers sequences of the corresponding SNP markers of Hordeum vulgare nuclear DNA, as well as the size of amplified loci of discriminated alleles [46]. The procedure of gene identification is carried out by the Amplification Refractory Mutation System (ARSM-PCR), followed by data interpretation by horizontal gel electrophoresis or by high resolution melting curves (HRM) analysis on PCR platforms in real time. It should be mentioned that we selected five SNP-markers (out of nine described by Chiapparino et al. [46] as potentially suitable for beer DNA authentication due to generation of relatively small allele-specific PCR products, whose size was not more than 200 bp (Table 4).
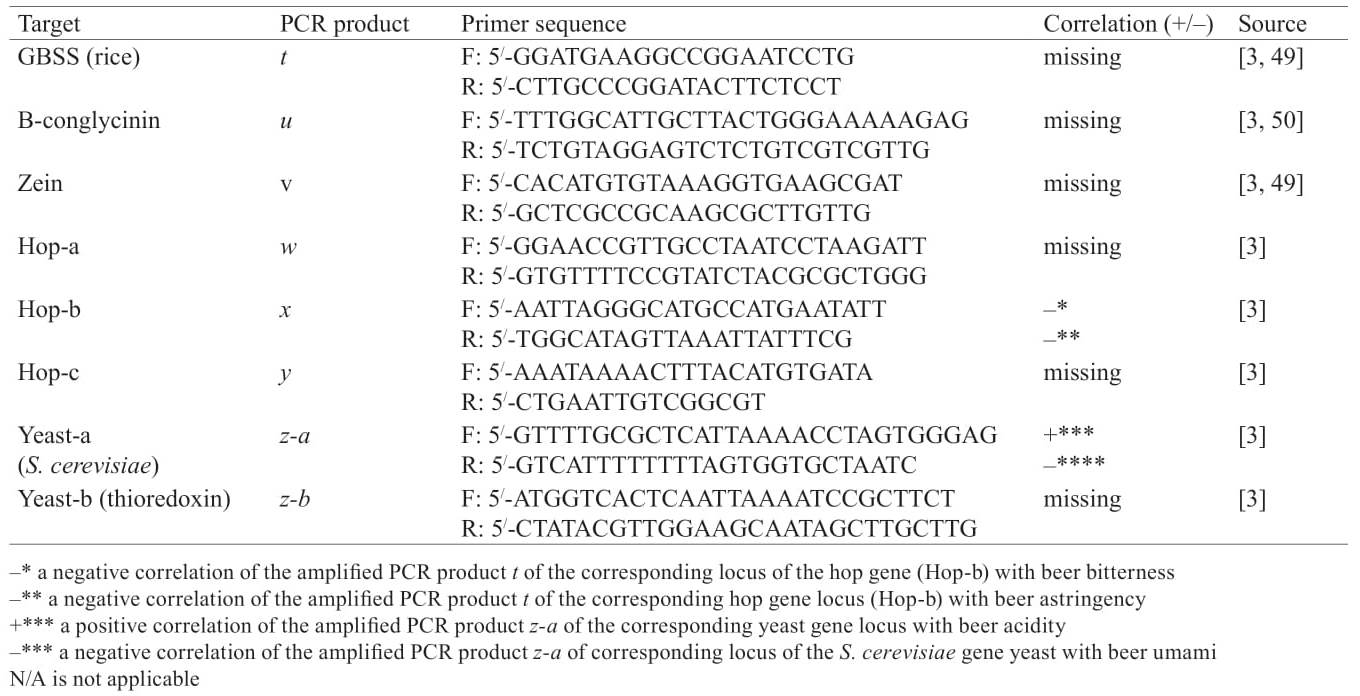
The detection of brewing barley substitutes in beer, which is often used as a cheap source of starch, makes it possible to evaluate the products sold for qualitative, quantitative, information and complex falsification. Table 5 demonstrates primer sequences targeting genetic targets used in the detection of brewing barley substitutes in beer, such as granulebound starch synthase of rice, β-conglycinin of soya, and zein of maize [49, 50]. Nevertheless, other PCR systems developed for the identification of cereals in food products can also be suitable for beer DNA authentication [51].
The effect of hops and yeast on beer quality is well-known. Thus, hop has a bactericidal effect on beer as well as provides its bitterness, aroma and foam stability [52]. Yeast is used in beer fermentation and impacts its character and taste [53]. Table 5 also presents sets of primers which initiate the amplification of specific PCR products of the corresponding loci of hops and yeast genes. They also allow the identifying or differentiating of commercial beer samples [3]. In addition, a negative correlation of the amplified PCR product t of the corresponding locus of the hop gene (Hop-b) with beer bitterness and astringency was revealed. The amplified PCR product z-a of the corresponding locus of the yeast gene S. cerevisiae showed a positive correlation with beer acidity and a negative correlation with beer umami [3]. Taking into account the rapid development of genomic and bioinformation technologies, metagenomic analysis, which allows determining yeast species diversity in beer samples without microorganisms allocating and cultivating, is one of the promising approaches to beer DNA authentication [54, 55].
ВЫВОДЫ
Analysis of scientific and methodical approaches to extraction of residual quantities of nucleic acids of beer raw materials and beer DNA-authentication indicates the applicability of molecular and genetic analysis in detecting counterfeit and falsified brewery products. The use of DNA technologies helps determine the authenticity and origin of the brewery industry products. Molecular labelling systems suitable for identification of Hordeum vulgare barley malt, or its substitutes, as well as hops and yeast, can ensure traceability of the product life cycle. Systematic data on correlation of amplified DNA targets with beer quality indicators can be of practical importance when choosing raw materials for brewery production.
КОНФЛИКТ ИНТЕРЕСОВ
The authors state that there is no conflict of interest.
ФИНАНСИРОВАНИЕ
The study was supported by the section of storage and processing of agricultural products of the Department of Agricultural Sciences of the Russian Academy of Sciences and Federal scientific centre ‘Food Systems’ of the Russian Academy of Sciences.
СПИСОК ЛИТЕРАТУРЫ
- Oganesyants LA, Khurshudyan SA, Galstyan AG. Monitoring kachestva pishchevykh produktov – bazovyy ehlement strategii [Food quality monitoring is a strategy key part]. Production Quality Control. 2018;(4):56–59. (In Russ.).
- Lachenmeier DW. Advances in the detection of the adulteration of alcoholic beverages including unrecorded alcohol. In: Downey G, editor. Advances in Food Authenticity Testing. Amsterdam: Woodhead Publishing; 2016. pp. 565–584. DOI: https://doi.org/10.1016/B978-0-08-100220-9.00021-7.
- Nakamura S, Tsushima R, Ohtsubo K. A Novel Method for the Preparation of Template DNA for PCR from Beer to Detect Materials and to Develop DNA Markers to Evaluate the Quality of Beer. Bioscience Biotechnology and Biochemistry. 2013;77(4):820–831. DOI: https://doi.org/10.1271/bbb.120969.
- Kuballa T, Brunner TS, Thongpanchang T, Walch SG, Lachenmeier DW. Application of NMR for authentication of honey, beer and spices. Current Opinion in Food Science. 2018;19:57–62. DOI: https://doi.org/10.1016/j.cofs.2018.01.007.
- Nakamura S, Haraguchi K, Mitani N, Ohtsubo K. Novel Preparation Method of Template DNAs from Eine for PCR To Differentiate Grape (Vitis vinifera L.) Cultivar. Journal of Agricultural and Food Chemistry. 2007;55(25):10388–10395. DOI:https://doi.org/10.1021/jf072407u.
- Ohtsubo K, Suzuki K, Haraguchi K, Nakamura S. Novel method for preparation of the template DNA and selection of primers to differentiate the material rice cultivars of rice wine by PCR. Journal of Biochemical and Biophysical Methods. 2008;70(6):1020–1028. DOI: https://doi.org/10.1016/j.jbbm.2007.07.001.
- Kim CS, Lee CH, Shin JS, Chung YS, Hyung NI. A simple and Rapid Method for Isolation of High Quality Genomic DNA from Fruit Trees and Conifers Using PVP. Nucleic Acids Research. 1997;25(5):1085–1086. DOI: https://doi.org/10.1093/nar/25.5.1085.
- Koonjul PK, Brandt WF, Farrant JM, Lindsey GG. Inclusion of polyvinylpyrrolidone in the polymerase chain reaction reverses the inhibitory effects of polyphenolic contamination of RNA. Nucleic Acids Research. 1999;27(3):915–916. DOI: https://doi.org/10.1093/nar/27.3.915.
- Juvonen R, Haikara A. Amplification Facilitators and Pre-Processing Methods for PCR Detection of Strictly Anaerobic Beer-Spoilage Bacteria of the Class Clostridia in Brewery Samples. Journal of the Institute of Brewing. 2009;115(3):167–176. DOI: https://doi.org/10.1002/j.2050-0416.2009.tb00365.x.
- Catalano V, Moreno-Sanz P, Lorenzi S, Grando MS. Experimental Review of DNA-Based Methods for Wine Traceability and Development of a Single-Nucleotide Polymorphism (SNP) Genotyping Assay for Quantitative Varietal Authentication. Journal of Agricultural and Food Chemistry. 2016;64(37):6969–6984. DOI: https://doi.org/10.1021/acs.jafc.6b02560.
- Pulido A, Bakos F, Devic M, Barnabas B, Olmedilla A. HvPG1 and ECA1: two genes activated transcriptionally in the transition of barley microspores from the gametophytic to the embryogenic pathway. Plant Cell Reports. 2009;28(4):551–559. DOI: https://doi.org/10.1007/s00299-008-0662-2.
- Pomortsev AA, Martynov SP, Lialina EV. Hordein Locus Polymorphism in Near Eastern Local Populations of Cultivated Barley (Hordeum vulgare L.). Genetika. 2008;44(6):815–828. (In Russ.).
- Lyalina EV, Boldyrev SV, Pomortsev AA. Current state of the genetic polymorphism in spring barley (Hordeum vulgare L.) from Russia assessed by the alleles of hordein-coding loci. Genetika. 2016;52(6):650–663. (In Russ.).
- Yamaguchi O, Baba T, Furusho M. Relationship between genotype of hordein and malting quality in Japanese barley. Breeding Science. 1998;48(3):309–314.
- Echart-Almeida C, Cavalli-Molina S. Hordein polypeptide patterns in relation to malting quality in Brazilian barley varieties. Pesquisa Agropecuaria Brasileira. 2001;36(2):211–217. DOI: https://doi.org/10.1590/s0100-204x2001000200001.
- Nakamura S, Suzuki K, Haraguchi K, Yoza K, Okunishi T, Matsui T, et al. Identification of domestic glutinous rice cultivars by the PCR method using grains of 18 typical glutinous rice cultivars as sample and development of technology for detection of different kind grain incorporation in glutinous rice processed foodstuffs. Nippon Nogeikagaku Kaishi-Journal of the Japan Society for Bioscience Biotechnology and Agrochemistry. 2004;78(10):984–993. DOI: https://doi.org/10.1271/nogeikagaku1924.78.984.
- Brandt A, Montembault A, Cameronmills V, Rasmussen SK. Primary structure of A B1 hordein gene from barley. Carlsberg Research Communications. 1985;50(6):333–345. DOI: https://doi.org/10.1007/bf02907156.
- Washington JM, Box A, Barr AR. Developing waxy barley cultivars for food, feed and malt. International Symposium ‘Barley Genetics’; 2000; Adelaide. Adelaide: The University of Adelaide; 2000. pp. 303–306.
- Clarke B, Liang R, Morell MK, Bird AR, Jenkins CLD, Li Z. Gene expression in a starch synthase IIa mutant of barley: changes in the level of gene transcription and grain composition. Functional & Integrative Genomics. 2008;8(3):211–221. DOI: https://doi.org/10.1007/s10142-007-0070-7.
- Rohde W, Becker D, Salamini F. Structural-analysis of the waxy locus from Hordeum vulgare. Nucleic Acids Research. 1988;16(14):7185–7186. DOI:https://doi.org/10.1093/nar/16.14.7185.
- Nakamura S, Machida K, Ohtsubo K. Search for Cell-Wall-Degrading Enzymes of World-Wide Rice Grains by PCR and Their Effects on the Palatability of Rice. Bioscience Biotechnology and Biochemistry. 2012;76(9):1645–1654. DOI: https://doi.org/10.1271/bbb.120147.
- Rasmussen SK, Klausen J, Hejgaard J, Svensson B, Svendsen I. Primary structure of the plant serpin BSZ7 having the capacity of chymotrypsin inhibition. Biochimica Et Biophysica Acta – Protein Structure and Molecular Enzymology. 1996;1297(2):127–130. DOI: https://doi.org/10.1016/s0167-4838(96)00115-x.
- Iimure T, Takoi K, Kaneko T, Kihara M, Hayashi K, Ito K, et al. Novel Prediction Method of Beer Foam Stability Using Protein Z, Barley Dimeric α-Amylase Inhibitor-1 (BDAI-1) and Yeast Thioredoxin. Journal of Agricultural and Food Chemistry. 2008;56(18):8664–8671 DOI: https://doi.org/10.1021/jf801184k.
- Niu CT, Han YP, Wang JJ, Zheng FY, Liu CF, Li YX, et al. Malt derived proteins: Effect of protein Z on beer foam stability. Food Bioscience. 2018;25:21–27. DOI: https://doi.org/10.1016/j.fbio.2018.07.003.
- Iimure T, Kihara M, Ichikawa S, Ito K, Takeda K, Sato K. Development of DNA markers associated with beer foam stability for barley breeding. Theoretical and Applied Genetics. 2011;122(1):199–210. DOI: https://doi.org/10.1007/s00122-010-1436-0.
- Knox CA, Sonthayanon B, Chandra GR, Muthukrishnan S. Structure and organization of two divergent α-amylase genes from barley. Plant Molecular Biology. 1987;9(1):3–17. DOI: https://doi.org/10.1007/BF00017982.
- Paris M, Jones MGK, Eglinton JK. Genotyping single nucleotide polymorphisms for selection of barley β-amylase alleles. Plant Molecular Biology Reporter. 2002;20(2):149–159. DOI: https://doi.org/10.1007/BF02799430.
- Abbott MS, Fedele MJ. A DNA-based varietal identification procedure for hops leaf tissue. Journal of the Institute of Brewing. 1994;100(4):283–285. DOI: https://doi.org/10.1002/j.2050-0416.1994.tb00825.x.
- Wei FS, Wing RA, Wise RP. Genome Dynamics and Evolution of the Mla (Powdery Mildew) Resistance Locus in Barley. Plant Cell. 2002;14(8):1903–1917. DOI: https://doi.org/10.1105/tpc.002238.
- Hirota N, Kaneko T, Kuroda H, Kaneda H, Takashio M, Ito K, et al. Characterization of lipoxygenase-1 null mutants in barley. Theoretical and Applied Genetics. 2005;111(8):1580–1584. DOI: https://doi.org/10.1007/s00122-005-0088-y.
- Hirota N, Kuroda H, Takoi K, Kaneko T, Kaneda H, Yoshida I, et al. Brewing Performance of Malted Lipoxygenase-1 Null Barley and Effect on the Flavor Stability of Beer. Cereal Chemistry. 2006;83(3):250–254. DOI: https://doi.org/10.1094/CC-83-0250.
- Yu JH, Huang SX, Dong JJ, Fan W, Huang SL, Liu J, et al. The influence of LOX-less barley malt on the flavour stability of wort and beer. Journal of the Institute of Brewing. 2014;120(2):93–98. DOI: https://doi.org/10.1002/jib.122.
- Oozeki M, Sotome T, Haruyama N, Yamaguchi M, Watanabe H, Okiyama T, et al. The two-row malting barley cultivar ‘New Sachiho Golden’ with null lipoxygenase-1 improves flavor stability in beer and was developed by marker assisted selection. Breeding Science. 2017;67(2):165–171. DOI: https://doi.org/10.1270/jsbbs.16104.
- Nagamine T, Amagai M, Ikeda TM, Oozeki M, Haruyama N, Kato T, et al. Development and evaluation of DNA markers for Japanese malting barley [Hordeum vulgare] breeding. Bulletin of the Tochigi Prefectural Agricultural Experiment Station (Japan). 2008;59:45–54.
- van Mechelen JR, Smits M, Douma AC, Rouster J, Cameronmills V, Heidekamp F, et al. Primary structure of a lipoxygenase from barley-grain as deduced from its CDNA sequence. Biochimica Et Biophysica Acta – Lipids and Lipid Metabolism. 1995;1254(2):221–225. DOI: https://doi.org/10.1016/0005-2760(94)00231-m.
- Perovic D, Kopahnke D, Habekuss A, Ordon F, Serflina A. Marker-Based Harnessing of genetic diversity to improve resistance of barley to fungal and viral disease. In: Miedaner T, Korzun V, editors. Applications of Genetic and Genomic Research in Cereals. Woodhead Publishing; 2018. pp. 137–164. DOI: https://doi.org/10.1016/B978-0-08-102163-7.00007-7.
- Rodriguez-Palenzuela P, Royo J, Gomez L, Sanchez-Monge R, Salcedo G, Molina-Cano JL, et al. The gene for trypsin-inhibitor CMe is regulated in trans by the lys 3a locus in the endosperm of barley (Hordeum Vulgare L). Molecular & General Genetics. 1989;219(3):474–479. DOI: https://doi.org/10.1007/bf00259622.
- Diaz I, Royo J, Oconnor A, Carbonero P. The promoter of the gene Itr1 from barley confers a different tissue-specificity in transgenic tobacco. Molecular and General Genetics. 1995;248(5):592–598. DOI: https://doi.org/10.1007/bf02423455.
- Henry RJ, Cowe IA. Factors influencing the hardness (milling energy) and malting quality of barley. Journal of the Institute of Brewing. 1990;96(3):135–136. DOI: https://doi.org/10.1002/j.2050-0416.1990.tb01024.x.
- Tonooka T, Aoki E, Yoshioka T, Taketa S. A novel mutant gene for (1-3, 1-4)-β-D-glucanless grain on barley (Hordeum vulgare L.) chromosome 7H. Breeding Science. 2009;59(1):47–54. DOI: https://doi.org/10.1270/jsbbs.59.47.
- Burton RA, Jobling SA, Harvey AJ, Shirley NJ, Mather DE, Bacic A, et al. The Genetics and Transcriptional Profiles of the Cellulose Synthase-Like Hvcslf Gene Family in Barley. Plant Physiology. 2008;146(4):1821–1833. DOI: https://doi.org/10.1104/pp.107.114694.
- Mei L, Ping J, Wang D, Zhang Z, Luo S, Yang M, et al. Malt genotypic screening of polymorphism information content (PIC) of PCR-based marker in barley, based on physiological traits. Molecular Biology. 2012;1(1):101–106. DOI: https://doi.org/10.4172/2168-9547.1000101.
- Lakhneko OR, Morgun BV, Kalendar RM, Stepanenko AI, Troianovska AV, Rybalka OI. SSR analysis in the study of genetic diversity and similarity of barley cultivars. Biotechnologia Acta. 2016;9(3):61–68. DOI: https://doi.org/10.15407/biotech9.03.061.
- Jo WS, Kim HY, Kim KM. Development and characterization of polymorphic EST based SSR markers in barley (Hordeum vulgare). 3 Biotech. 2017;7. DOI: https://doi.org/10.1007/s13205-017-0899-y.
- Tomka M, Urminska D, Canapek M, Galova Z. Potential of selected SSR markers for identification of malting barley genotypes. Journal of Microbiology, Biotechnology and Food Sciences. 2017;6(6):1276–1279. DOI: https://doi.org/10.15414/jmbfs.2017.6.6.1276-1279.
- Chiapparino E, Lee D, Donini P. Genotyping single nucleotide polymorphisms in barley by tetra-primer ARMS-PCR. Genome. 2004;47(2):414–420. DOI: https://doi.org/10.1139/g03-130.
- Tabone T, Mather DE, Hayden MJ. Temperature Switch PCR (TSP): Robust assay design for reliable amplification and genotyping of SNPs. Bmc Genomics. 2009;10:14. DOI: https://doi.org/10.1186/1471-2164-10-580.
- Hayden MJ, Tabone T, Mather DE. Development and assessment of simple PCR markers for SNP genotyping in barley. Theoretical and Applied Genetics. 2009;119(5):939–951. DOI: https://doi.org/10.1007/s00122-009-1101-7.
- Ohtsubo K, Nakamura S, Yoza K, Shishido K. Identification of glutinous rice cultivars using rice cake as samples by the PCR method. Journal of the Japanese Society for Food Science and Technology-Nippon Shokuhin Kagaku Kogaku Kaishi. 2001;48(4):306–310.
- Tsukada Y, Kitamura K, Harada K, Kaizuma N. Genetic Analysis of Subunits of Two Major Storage Proteins (β-Conglycinin and Glycinin) in Soybean Seeds. Japanese Journal of Breeding. 1986;36(4):390–400. DOI: https://doi.org/10.1270/jsbbs1951.36.390.
- Silletti S, Morello L, Gavazzi F, Giani S, Braglia L, Breviario D. Untargeted DNA-based methods for the authentication of wheat species and related cereals in food products. Food Chemistry. 2019;271:410–418. DOI: https://doi.org/10.1016/j.foodchem.2018.07.178.
- Kovacevic M, Kac M. Solid-phase microextraction of hops volatiles - Potential use for determination and verification of hops varieties. Journal of Chromatography A. 2001;918(1):159–167. DOI: https://doi.org/10.1016/s0021-9673(01)00719-1.
- Naumov GI, Naumova ES, Lantto RA, Louis EJ, Korhola M. Genetic homology between Saccharomyces cerevisiae and its sibling species S. paradoxus and S. bayanus: Electrophoretic karyotypes. Yeast. 1992;8( 8):599–612. DOI: https://doi.org/10.1002/yea.320080804.
- Sobel J, Henry L, Rotman N, Rando G. BeerDeCoded: the open beer metagenome project. F1000Res. 2017;6:1676. DOI: https://doi.org/10.12688/f1000research.12564.2.
- Batut B, Gravouil K, Defois C, Hiltemann S, Brugere JF, Peyretaillade E, et al. ASaiM: a Galaxy-based framework to analyze microbiota data. Gigascience. 2018;7(6). DOI: https://doi.org/10.1093/gigascience/giy057.